It’s called bird flu for a reason. Particular characteristics about the influenza virus known as H5N1 allow it to primarily affect avifauna, though in some worrying cases the disease has been passed to humans.
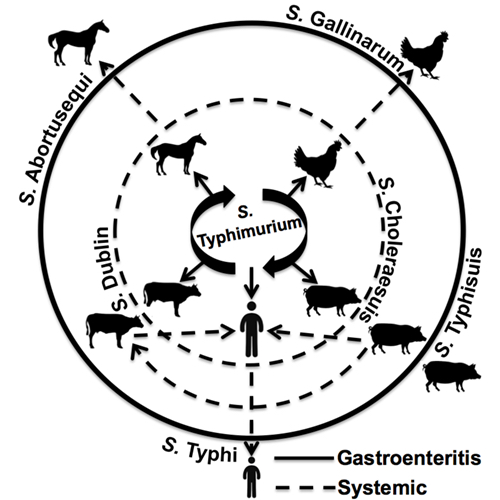
Similarly, many strains, known as serovars, of Salmonella bacteria are specific to certain types of animals. Some infect cows, others poultry and still others affect primarily humans.
Now, research by a team led by University of Pennsylvania scientists has shown, using genomic techniques, that slight variations in the coding sequence of proteins that bind Salmonella to host cells can determine what type of animal a particular strain infects.
The work appears in Nature Communications.
 “In Salmonella, we knew that many serovars are specific to one host; we didn’t know why, but we knew that they are,” said Dieter Schifferli, senior author on the paper and a professor of microbiology in Penn’s School of Veterinary Medicine. “In this work, we found strong associations between different serovars’ adhesin molecules and their preferred hosts, relationships that we then confirmed with work in the lab.”
“In Salmonella, we knew that many serovars are specific to one host; we didn’t know why, but we knew that they are,” said Dieter Schifferli, senior author on the paper and a professor of microbiology in Penn’s School of Veterinary Medicine. “In this work, we found strong associations between different serovars’ adhesin molecules and their preferred hosts, relationships that we then confirmed with work in the lab.”
The research relies on what are known as genome-wide association studies, or GWAS, in which the genomes of various strains of Salmonella were partially sequenced and then compared, looking at key characteristics. This work made use of an enormous library of Salmonella maintained by Penn Vet. It contains thousands of different strains, as well as samples from the United States Centers for Disease Control and Prevention, U.S. Food and Drug Administration, Pennsylvania Department of Health and Institut Pasteur.
The scientists first focused their analyses on Salmonella enterica serovar Typhimurium, which is a leading cause of food poisoning but for which the molecular basis for host preferences is still a mystery.
The genomic analyses identified single nucleotide polymorphisms, or SNPs, which are variations in the nucleotide sequence of DNA. The scientists found a relatively large number of SNPs, particularly those that result in the production of a different amino acid, in genes coding for Salmonella surface proteins or secreted factors.
 “We saw this huge variation in proteins on the surface of bacteria or in secretions, which are really the first lines of interaction with the host,” Schifferli said. “If there was so much variation, it suggests it must be linked to something important.”
“We saw this huge variation in proteins on the surface of bacteria or in secretions, which are really the first lines of interaction with the host,” Schifferli said. “If there was so much variation, it suggests it must be linked to something important.”
Indeed, the researchers’ analysis showed that different host species tended to share patterns of these so-called non-synonymous SNPs, which create different protein sequences and structures.
Following these initial findings, the research team focused on genes for adhesin proteins, which play a key role in the interaction between bacteria and host. Analyzing 15 genes in 580 strains of Typhimurium, they found a high degree of variation and evidence of positive selection and strong evidence that the variation was associated with the particular strains’ host specificity.
With this statistical support in hand, the researchers went into the lab to test their findings. They chose to closely examine the adhesive properties of the protein FimH encoded by the gene fimH, which had shown the most variation in their previous analyses, a total of 17 variants, for their experiments. Selecting one variant that had been associated with human samples and another that had been associated with bovine samples, they introduced the variants into Escherichia coli and then tested the bacteria’s resulting binding affinity to cultured cells of either bovine or human origin.
 Though the only difference between the two FimH variants was a single amino acid, they found that the bovine-associated FimH indeed bound preferentially to all bovine cells compared to the human-associated FimH. Selectively altering this one amino acid in a human-associated strain of Salmonella effectively reduced the ability of that strain to bind to human cells and increased its affinity for bovine cells.
Though the only difference between the two FimH variants was a single amino acid, they found that the bovine-associated FimH indeed bound preferentially to all bovine cells compared to the human-associated FimH. Selectively altering this one amino acid in a human-associated strain of Salmonella effectively reduced the ability of that strain to bind to human cells and increased its affinity for bovine cells.
“Manipulating bacteria in this way allowed us to show a cause and effect relationship between a SNP in an adhesin gene and the bacteria’s host specificity,” Schifferli said.
Further studies expanded this look to more serovars beyond Typhimurium, finding similar patterns of host-specific variation in the FimH protein.
A final set of experiments, again in the lab, expressed a variety of FimH variants from multiple serovars in E. coli, and tested their binding potential to porcine, human, bovine and chicken cells in culture. They found distinct species binding preferences for many of the FimH variants, confirming the host-specific associations they had seen in the in silico, or computational analyses.
Schifferli and colleagues plan to do similar genomic association studies to determine the genetic differences that may separate a strain of Salmonella that causes a brief gastrointestinal illness from one that cause a major systemic disease.
“The power of bacterial genomic association studies is that they can guide your work in the lab,” he said.
Co-authors on the paper included Penn Vet’s Min Yue, Xiangan Han, Leon De Masi, Chunhong Zhu, Xun Ma, Junjie Zhang, Renwei Wu and Shelley C. Rankin; Dustin Brisson of Penn’s Department of Biology; Nancy R. Zhang of Penn’s Wharton School; San Diego State University’s Robert Schmieder and Robert A. Edwards, also of Argonne National Laboratory; South Dakota State University’s Radhey S. Kaushik; the Pennsylvania Department of Health’s George P. Fraser; the FDA’s Shaohua Zhao and Patrick F. McDermott; the Institut Pasteur’s François-Xavier Weill; University of Liège’s Jacques G. Mainil; and the University of Maryland’s Cesar Arze and W. Florian Fricke.
The study was supported by the National Institutes of Health, the U.S. Department of Agriculture, the University of Pennsylvania Veterinary Center for Infectious Disease, Penn Vet’s Center for Host-Microbial Interactions and the China Scholarship Council.